How do we track Covid-19 variants?
Read our explainer on how genomic surveillance can help us track and contain Covid-19 variants.
From Omicron to Pirola, new Covid-19 variants are still appearing. But what are variants? We asked Titus Divala, Epidemics Research Lead, to explain how variants happen, what they mean for treatments and vaccines, and how we can prevent them.
Wellcome

Interim Head of Epidemics & Epidemiology, Wellcome

Interim Head of Epidemics & Epidemiology, Wellcome
This article was originally published on 29 November 2021, and updated on 18 October 2023.
Although the terms viral mutants, variants and strains are often used interchangeably, they generally hold different meanings.
To spread, a virus needs to infect a host, replicate and produce lots of copies of itself.
When a virus replicates, it doesn’t always manage to produce an exact copy of itself. This means that, over time, the virus may start to differ slightly in terms of its genetic sequence. Any change to the viral genetic sequence during this process is known as a mutation, and viruses with new mutations are sometimes called variants. Variants can differ by one or multiple mutations.
When a new variant has different functional properties to the original virus and becomes established in a population, it is sometimes referred to as a new strain of the virus.
In short, all strains are variants, but not all variants are strains.
All viruses mutate, but most mutations are harmless and do not affect the properties of the virus.
However, some mutations can give the virus an advantage, such as making it more transmissible (able to spread more easily) or better at evading the immune system.
These mutations can lead to the emergence of new variants of the virus, which can affect:
Pressing play on the video above will set a third-party cookie. Please read our cookie policy for more information.
Sir Jeremy Farrar explains why viruses mutate and the risks mutations pose to vaccines and treatments.
Why does the Covid-19 virus mutate?
Anna Mouser, former Policy and Advocacy Lead, Vaccines: I wondered if you would describe for a layperson, a non-scientist, this issue of the virus evolving. How would you explain that to somebody without Biology A-Level or similar?
Sir Jeremy Farrar, OBE, Director of the Wellcome Trust (2013-2023): RNA viruses change a lot. HIV viruses are RNA viruses and influenza viruses are RNA viruses, and they mutate. They mutate randomly, there's no thinking behind it. It just happens. And then those mutations which give the virus an advantage, it can more easily go from me to you. It can transmit easier, it can pass to other people. Those viruses will become the dominant strains in a community. We've seen that in the UK, South Africa as well, where a mutation happened. It gave the virus an advantage. It was easier to transmit from one person to another and it became the dominant strain. What is more worrying is the fact that we now have people that have been infected and we're now starting to vaccinate people. Therefore, the virus that mutates in a way that escapes those vaccines or infections will also give those viruses an advantage and they will become the dominant strains. The best way of preventing that is to reduce the amount of viruses in the world, reduce the chances of mutations happening and reduce the chance of that variant having an advantage or becoming dominant. And the best way of doing that is to reduce transmission, make the vaccines available globally, and prevent those variants from happening. It's much easier to prevent than it is to respond to new variants coming into the population of viruses in the world.
There have been many mutations of the virus (SARS-CoV-2) that cause Covid-19 since it was first identified.
Not all variants are a cause for concern. Public health authorities determine which Covid-19 variants to worry about by evaluating research on the variant's potential implications, guided by the World Health Organization’s classification framework.
The World Health Organization classifies variants based on their potential impact on transmissibility, disease severity, immune evasion, and diagnostics and therapeutics.
Each new Covid-19 variant is assigned a Greek letter in the order they are identified. This naming style brings simplicity and allows the public to remember and keep track.
You may have heard of Alpha, Beta, Gamma, Delta and Omicron. However, there are many less popular Covid-19 variants, such as Epsilon, Zeta, Eta, Theta, Iota, Kappa, Lambda and Mu, which never became variants of concern. In March 2023, the World Health Organization restricted the Greek letter naming to only variants of concern.
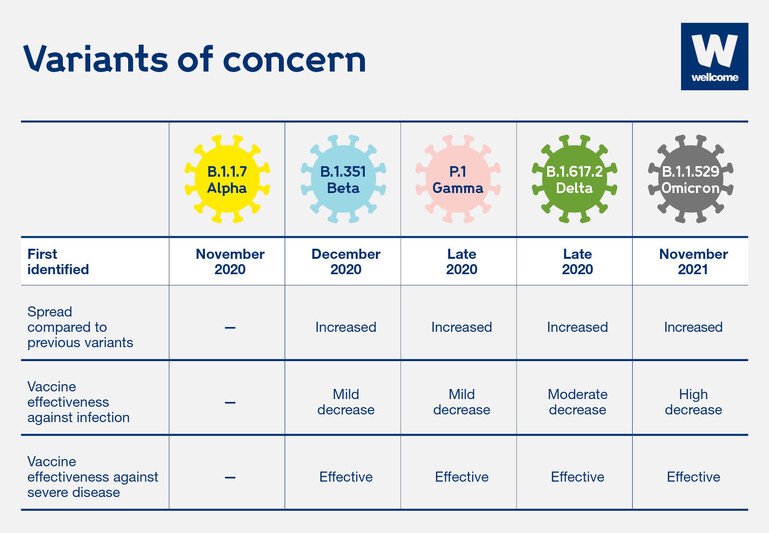
A table of key Covid-19 variants (Alpha, Beta, Gamma, Delta and Omicron), when they emerged and why they are of concern.
Source: https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/
Wellcome
Omicron is the most dominant of the Covid-19 variants, and several subvariants have emerged since it was first identified, including BA.2.86 (nicknamed Pirola) and EG.5.1.
Stay updated with the latest Covid-19 variants via the World Health Organization’s tracking database.
Pressing play on the video above will set a third-party cookie. Please read our cookie policy for more information.
How Covid-19 variants happen
All viruses naturally mutate. Sometimes a virus gets lucky and mutates in a way that helps it survive and spread more easily.
The virus that causes Covid-19 is no exception.
New variants have been spreading across the world.
They seem to transmit more easily. This could mean more people getting infected and more pressure on already stretched health services.
But we don't know yet how well existing treatments and vaccines will work against known and future variants.
To give the virus fewer opportunities to mutate, we need to cut transmission, increase the pace of research into Covid-19 treatments and vaccines and make them available fairly across the world.
Scientists need more funding for Covid-19 research.
Variants have the potential to make current treatments and vaccines less effective. This is due to individual mutations that may reduce the virus’s sensitivity and vulnerability to antibodies (part of our immune response).
For example, many Covid-19 vaccines target the spike protein that sits on the virus's surface and comes into contact with our immune system. Therefore, a virus that accumulates numerous mutations in the spike protein may be able to evade natural or vaccine-induced immunity.
There is ongoing research to test the efficacy of existing Covid-19 vaccine candidates against a range of variants. For example, scientists were initially concerned about the BA.2.86 variant because of its high number of mutations and potential to evade existing immunity. However, evidence as of September 2023 suggests that the public health risk is low.
To ensure that vaccines, diagnostics and treatments stay effective, researchers and developers respond to new variants by updating their vaccines, diagnostics and treatments as the virus changes.
The influenza virus, which causes flu, is known to be a frequently mutating virus. Every flu season, we see several different influenza variants in circulation, which means that existing vaccines need to be updated so they are effective against the new variants.
This is why it is important to continue to invest in the development of a variety of vaccines and treatments to increase the chance of having tools that are effective against new variants.
Due to the potential for a mutation to change the properties of a virus, we need to monitor variants closely to determine if they pose a greater risk of transmission, severe disease or evasion of current interventions.
Since the 1980s, we’ve been monitoring and sequencing influenza (flu) virus strains through global networks to enable adaptation of the flu vaccine every year. And now, the same techniques are being used to monitor and adapt to Covid-19 variants.
This is done through genetic sequencing of a virus and the collection of epidemiological and clinical data. Scientists monitor any changes to a virus’s genome and assess the variant's impact on diagnostics, vaccines and therapeutics. When concerning changes are identified, they are widely reported, enabling the adaptation of public health interventions and policies.

Read our explainer on how genomic surveillance can help us track and contain Covid-19 variants.
Public health authorities and researchers monitor Covid-19 variants by collecting data on their prevalence, geographic distribution, potential for international spread, and the impact of variants on diagnostics, therapeutics and vaccines.
However, the capacity to deliver this virus monitoring varies between countries. This is a concern because we know that viruses – such as Covid-19 – do not respect geographical borders, and new variants spread across countries. The lack of international genomic surveillance also leads to gaps in data and poses the risk of new harmful variants emerging and spreading undetected.
Therefore, rapid sharing of genetic sequencing and relevant epidemiological data associated with Covid-19 is crucial. This data enables researchers and health authorities to determine if a new variant poses a greater risk of transmission, severe disease or evasion of current interventions. Collection of this data also helps target vaccines and therapeutics development and helps monitor their impact when they are introduced.
On the flip side, if an increase in cases is noticed somewhere that cannot be explained or linked to a super-spreader event (for example, a workplace outbreak), health authorities may review the genetic surveillance data to look for mutations that could have caused the increase.
The more virus that is in circulation, the more opportunity it has to replicate and mutate, potentially producing new variants of concern.
Suppressing the spread of a virus reduces the number of new variants, while also protecting populations from existing variants. To limit the spread, governments can: