This is how flooding can impact our health
Climate change is making the world more susceptible to extreme weather events like flooding, which can impact human health
Twenty-four research teams in 12 countries will receive a total of £22.7 million to develop new digital tools to respond to the emerging threat of climate-sensitive infectious diseases.
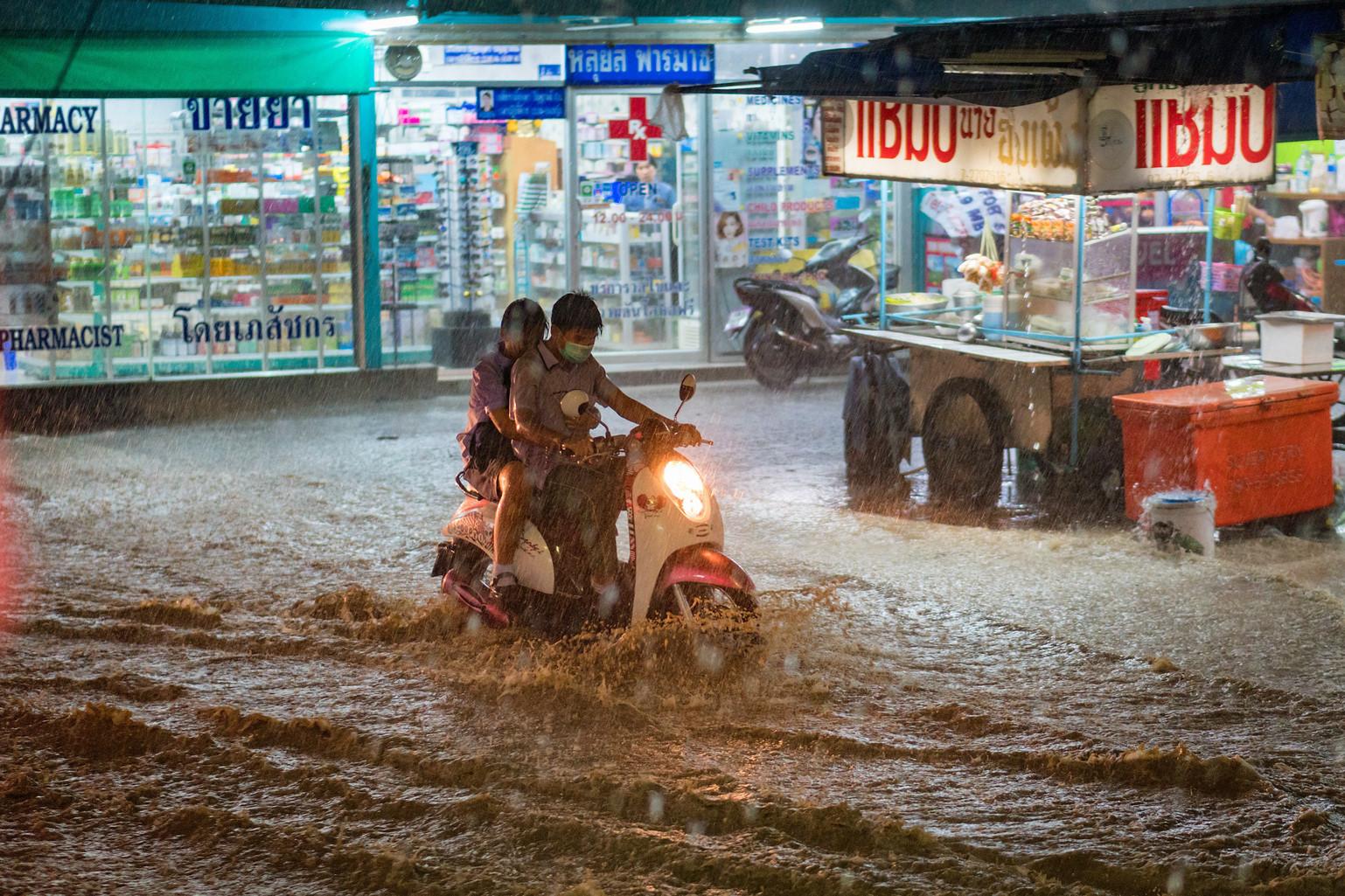

Qimono / Pixabay
Digital Content and Social Media Producer, Wellcome
Digital Content and Social Media Producer, Wellcome
We know climate change is having a profound effect on infectious diseases. Extreme weather events and warming temperatures are giving rise to infectious diseases and providing more opportunities for them to expand to new regions, putting the lives of billions of people at risk.
But what if we had early warning systems to predict the risk of disease outbreaks like cholera and dengue long before they happen? Or a tool that could estimate how changes to climate and land use will affect disease transmission? Or an app that could track the spread of diseases during natural disasters like floods?
If we knew more about when and where deadly disease outbreaks are likely to happen, policymakers could plan interventions to minimise their impact on communities that are most at risk.
Our new funding could be the next step in making that a reality.
Last year, we launched a funding call to support the development of new open-source technology – technology that is available to the public – to respond to the emerging threat of climate-sensitive infections and improve disease modelling.
The aim was to develop tools that use climate data to better understand and predict climate-sensitive infectious diseases, which could be used by policymakers and decision-makers to prepare and respond to potential outbreaks or changes in their distribution.
As part of this call, we’re now funding the development of 20 new tools across four categories:
Tools to help investigate the impacts of climate on infectious disease-related risks. These will help identify locations and times when the risk of disease transmission increases to prevent and control disease outbreaks.
Arboverse – a data-driven platform and disease modelling to anticipate and mitigate arbovirus emergence
Arboviruses are diseases transmitted to humans through the bite of an infected insect, such as mosquitoes and ticks. Led by researchers at the University of Texas, USA, Arboverse will create a user-friendly web platform that collates and processes data on arboviruses, vectors and environmental factors such as climate change, deforestation and human mobility – from a range of sources to give insight into the causes of arbovirus outbreaks.
WaterPath – future scenario toolkit for waterborne infectious disease modelling
Waterborne diseases include cholera, E. coli, AMR E. coli and schistosomiasis. Led by a team at Wageningen University, Netherlands, WaterPath will provide an open-source modelling toolkit to measure the impact of climate change and socio-economic development on the spread of waterborne pathogens and the disease risk they present. The toolkit will be based on an existing model, which will be extended to create projections based on climate data.
Climate-sensitive vector-borne disease intervention tools
Vector-borne diseases occur when the host (such as a human or an animal) is infected by a pathogen (such as a parasite). Pathogens can be passed from animal to animal, animal to human, or human to human through the infectious bite of different organisms such as mosquitoes, sandflies or ticks. Examples of vector-borne diseases include malaria, dengue, and yellow fever.
A team at Imperial College London, UK, will develop a flexible, user-friendly interface for policymakers and researchers to explore the impact of environmental change on the spread of vector-borne diseases. Users will be able to input disease projections and investigate – through simulation – the necessary public health measures to mitigate the effects of climate change.
Climate-driven models to predict future risk of arenaviruses
Arenaviruses are viruses normally found in rodents, and occasionally humans. The virus is transmitted to humans by the inhalation of the virus released from the urine, saliva or droppings of the infected rodent.
The team at One Health Institute at the University of California, Davis, USA, will develop a semi-automated data collection tool for arenaviruses, including data on rodent populations, human cases and trends in annual and extreme temperature and rainfall levels. Based on this data, models will be developed to understand future risks based on specific emission scenarios and local climate projections.
The impact of climate variability on infectious disease epidemic risks
A team at the University of Warwick, UK, will develop open-source software for estimating future epidemic risks in different locations. It will account for both climate change and natural climate variability such as the amount of sunlight and volcanic eruptions. The team will focus on vector-borne diseases, but the software could be adapted for other types of pathogens that are not vector-borne such as those that cause respiratory diseases.
Tools to investigate and predict the transmission risk of mosquito-borne diseases, with useful metrics to guide advanced resource allocation and intervention planning.
Digital technology for storm-related malaria response in Mozambique
Led by researchers at the University of Minnesota, USA, this project will develop models to identify areas in Mozambique where severe weather events will increase malaria risk. The team will also develop and pilot a software platform to predict areas at risk of malaria transmission after severe weather events, which could inform malaria control activities in those areas. The platform will be integrated into the malaria control and disaster management programmes already in place in Mozambique.
CLIMSEDIS – a climate-sensitive disease forecasting tool
A team at the University of Liverpool, UK, will investigate if the same climate factors drive multiple climate-sensitive diseases across various countries in the Horn of Africa. They will co-develop, pilot and test models to forecast changes in selected diseases with the National Ministries of Health and Agriculture and other relevant regional health and policy organisations.
The team will consider associations between climate and health that haven’t been investigated before, such as the relationship between climate-sensitive infectious diseases and the Indian Ocean Dipole (also known as the Indian Niño) – an irregular change in sea surface temperatures in the western Indian Ocean.
Mosqlimate
Researchers at Fundação Getulio Vargas, Brazil, will develop a tool to estimate how changes to climate and land use will affect patterns of transmission for multiple arboviral diseases such as Zika, dengue and Chikungunya.
The tool will use data on climate and diseases to generate a risk measure which can then be integrated into the Brazilian early warning system ‘Infodengue’, already commonly used by policymakers.
Another tool developed through the same project, OviCounter, will be used to help improve mosquito surveillance data.
HydroVec – a digital surveillance platform for areas under threat of extreme hydro-meteorological events
Led by a team at Liverpool School of Tropical Medicine, UK, HydroVec is a platform to predict the transmission risk of vector-borne diseases in areas vulnerable to extreme hydro-meteorological events, such as thunderstorms, hailstorms, tornados and floods.
The team will work closely with key governmental and humanitarian agencies in Malawi to design a sustainable digital platform that can be used to inform disease control policy and practice.
An open-source framework for Rift Valley Fever forecasting
A team of researchers from EcoHealth Alliance, USA, will develop a tool to predict changes in incidences of Rift Valley Fever – a viral disease most seen in domesticated animals in sub-Saharan Africa.
The tool will integrate data on African climate, population, livestock and vegetation. Local and regional groups will be able to input data specific to their area to use the tool to tailor preventive and control measures to their local conditions.
DART (Dengue Advanced Readiness Tools) – an integrated digital system for dengue outbreak prediction and monitoring
Researchers at the University of Oxford, UK, and Oxford University Clinical Research Unit, Vietnam, will develop an automated forecasting system for dengue that integrates datasets from across Hanoi and Ho Chi Minh. The team will build a mobile and desktop application to support predictions of dengue within cities. The application will integrate weather and disease forecasts to improve understanding of the relationship between the two.
Climate-driven vector-borne disease risk assessment
A team of researchers from The Cyprus Institute, Cyprus, will build a platform that organises and combines environmental and climate data and up-to-date mosquito surveillance data. The data will be combined with large-scale models to predict changes in the number of mosquitoes in the area and disease outbreaks in eastern Europe and central Asia. The team will create tutorials and educational tools for public health specialists.
Tools to predict the transmission risk of climate-sensitive infectious diseases that are not mosquito-borne – an area that is lacking research – to inform interventions. This includes respiratory diseases (like asthma), foodborne diseases (like Salmonella infection) and waterborne diseases (like cholera).
Chol-Out-EWS – an open-source software application for community-level cholera outbreak risk prediction and feedback through early warning systems
Led by a team at the International Centre for Diarrhoeal Disease Research, Bangladesh, Chol-Out-EWS will be an open-source software to model and predict the risk of cholera outbreaks in Bangladesh. The project will also involve communicating those risks through a user-friendly early warning system for policymakers and support timely control of infection transmission and disease management in identified hotspots.
A web app for accessible, reproducible, multi-scale regression models for mapping climate-driven infectious diseases
A team at the University of Leicester, UK, will develop an online app which will use health and environmental data to make spatially refined predictions of zoonotic and vector-borne diseases in policy-relevant ways. The tool will be co-designed with public health bodies within countries with a high burden of these diseases, such as the Nigeria Center for Disease Control and Prevention, which is a named partner.
IDExtremes – a modelling tool to predict the probability of infectious disease outbreaks given compound extreme climatic events
Led by researchers at the Barcelona Supercomputing Center, Spain, this user-friendly open-source tool will be co-developed with stakeholders in Barbados, South Sudan, Nigeria, Brazil, Bangladesh and Nepal. Users will be able to input observed and forecasted meteorological indicators like droughts, floods or heatwaves. The tool will predict the probability of outbreaks of climate-sensitive infectious diseases (like malaria, dengue, and cholera) several months in advance.
Co-creation of a multi-pathogen predictive dashboard to improve health system responsiveness to climate-sensitive diseases in rural Madagascar
Co-created with the Ministry of Public Health of Madagascar and the non-governmental organisation Pivot, this prediction tool will be used to inform the health system and strengthen interventions against local diseases in a rural area of Madagascar. It will focus on malaria, diarrheal disease and acute respiratory infections, and will be integrated into an existing health information system.
FloDisMod – framework for flood and disease modelling
Led by researchers at the University of Texas, USA, this open-source software will model floods and storm surges – particularly those occurring in the Southern Texas border region – to better understand their impact on mosquito habitat, disease reporting, social vulnerability and health risks. The team will develop an early warning system for the Texas-Mexico border region, but it will be extended to soil-borne diseases as well as other geographies.
Tools that can support and apply to a variety of problem areas, for example, modelling, data acquisition and data visualisation.
Health RADAR: responsible access to data for analysis and research
Researchers at the University of Cape Town, South Africa, will develop an open-source, web-based platform that aims to provide a central data resource for disease and other relevant data (like health, climate, transmission, entomology, economic, demographic and intervention cost data) specific to climate-sensitive infectious disease modelling. The platform will focus on malaria transmission in Botswana, Eswatini, Namibia and South Africa.
WADIM (Water-Associated infectious Diseases in India: digital Management tools)
A team at Plymouth Marine Laboratory, UK, will develop tools to map sanitation conditions and disease spread in India. These tools can be rapidly updated, especially in the event of natural disasters, such as floods, or changes in environmental conditions, like monsoon rainfall. The data will be collected through an app designed for local citizens and primary health workers, allowing them to fill an important gap in information.
CliZod, compiling knowledge – a participatory database of modelling parameters for climate-sensitive zoonotic disease
Led by a team at Massey University, New Zealand, CliZod will be a searchable, interactive and participatory database of model frameworks for zoonotic diseases. Researchers will use the artificial intelligence approach of natural language processing to scan and pull out relevant information from the literature. This will then be inputted into a database to overcome the limitations of systematic reviews that are subject to human bias and time constraints.
In addition to the projects awarded through the funding call, we are supporting several other projects in this space. These include:
ARBOTHAI
Researchers at the Instituto de Salud Global de Barcelona, Spain, will adapt an existing platform for arboviral risk prediction in Spain for application in an endemic region – Thailand – so it can be used as an operational early warning tool at the province and city levels. The system predicts the risk of local outbreaks following the detection of infections while accounting for the local climate, vector population, urbanisation and socioeconomic factors. The project will also generate state-of-the-art projections of future dengue risk across Thailand using a series of emission and socioeconomic scenarios.
Dynamic infectious disease risk platform for decision-makers
A team at the University of California, USA, will create an online platform for robust data analytics to predict dengue incidence in Latin America while using data from Peru as evidence.
The platform will use environmental predictors and real-time climate factors to drive ensemble machine learning models, which is a machine learning approach to combine multiple other models in the prediction process. Users will be able to view and generate maps, highlight regional disease incidence hotspots, and download historical and predicted data directly from the tool’s primary user interface.
User-friendly digital prediction tool for dengue prevention
A team at the University of Queensland, Australia, will build a predictive dengue model that accurately predicts dengue risk at the district level two months in advance. An open-source, early warning system with a user-friendly interface aimed at local health practitioners will be developed to predict dengue outbreaks and aid early action. To our knowledge, this is the first ever project to evaluate the effectiveness of a climate-informed early warning system for a climate-sensitive infectious disease using a randomised control trial.
HARMONIZE
Led by a team at Barcelona Supercomputing Centre, Spain, this project will develop techniques for systematic error removal in climate data and collate more targeted information for specific areas, including low- and middle-income countries. The work will start with health ministries and research institutions in Peru, Brazil, Colombia and the Dominican Republic.
Overcoming barriers to implementation of CSID modelling tools and early warning systems
Wellcome will fund the development of a community of practice to help remove current barriers to the implementation of these tools and support better integration of climate-sensitive infectious disease research into national and international policy. The community of practice aims to improve collaboration between subject matter experts and software developers and communication between modellers and decision-makers.
Wellcome is also carrying out a global landscaping study to identify barriers to local uptake and implementation of climate-sensitive infectious disease early warning systems and identify pre-existing solutions.
As the world continues to warm, it will become more vulnerable to climate-sensitive infectious diseases that thrive in those conditions, including waterborne, foodborne and vector-borne diseases.
We’re already seeing the impacts on health.
In Malawi, over 1,000 people have died during the country’s largest outbreak of cholera in two decades. In Nepal, higher-altitude regions – where disease outbreaks are typically less common – are now experiencing higher rates of mosquito-borne dengue fever.

Climate change is making the world more susceptible to extreme weather events like flooding, which can impact human health
The data is warning us: up to 8.4 billion people could be at risk of diseases like dengue and malaria by the end of the century if emissions keep rising at current levels. And even in a best-case scenario, it would still be 6.1 billion people potentially threatened.
Tools that utilise data to predict and manage the spread of climate-sensitive diseases are becoming more crucial to minimise the risk of outbreaks.
Funding for these research projects follows a Wellcome-commissioned study that identified technology gaps in global climate-sensitive infectious disease preparedness.
The study found there to be only 37 fully developed and named tools in papers published in the last ten years – a sign that this area of research is not receiving enough investment and support.
Of the technologies that do exist, the majority are for vector-borne diseases, with much less for respiratory, foodborne and waterborne diseases. There’s also a lack of global representation, as most of the tools have been created by North American and European institutions.
In addition, only a quarter of the tools were found to be useful for decision-makers with limited real-world benefits for communities.
This funding award has been designed to fill these gaps.
“Digital technologies such as numerical models and early warning systems are some of the most powerful and useful tools available for understanding and potentially mitigating the impacts of climate on infectious diseases," said Felipe Colón, Wellcome's Technology Lead.
"The Digital Technologies Development Award was a great opportunity to improve the field of climate-sensitive infectious disease modelling that could inform actions to reduce the global burden of climate-sensitive infectious diseases.”

As climate change alters temperatures and weather patterns around the world, the risk of vector-borne diseases like dengue fever and Zika virus will increase. Here’s what that means for global health and what can be done to limit the damage.
While we already have some data to guide us, there is a lot we still don’t know about how infectious disease research and climate science intersect and how data can help inform policies and decision-making for practical action.
The tools we’re funding through this award provide an exciting opportunity to address some of those gaps. And in doing so, they can possibly transform preparations and responses to outbreaks of devastating diseases, potentially saving millions of lives.
“Digital technologies [...] are some of the most powerful and useful tools available for understanding and potentially mitigating the impacts of climate on infectious diseases.”

Felipe Colón
Technology Lead
Wellcome
Connect with Felipe:
We are funding other opportunities in this space that use data – particularly data centring the communities and people most impacted by climate change – to solve urgent health challenges.
Find out more about our Data for Science and Health programme.
We’re funding vital research into the impact climate change has on human health around the world, at national, regional and global levels. Explore our current funding call:
Advancing climate mitigation solutions with health co-benefits in low- and middle-income countries